Hybridization assay
A hybridization assay comprises any form of quantifiable hybridization i.e. the quantitative annealing of two complementary strands of nucleic acids, known as nucleic acid hybridization.[1]
Overview
[edit]In the context of biochemistry and drug development, a hybridization assay is a type of Ligand Binding Assay (LBA) used to quantify nucleic acids in biological matrices. Hybridization assays can be in solution or on a solid support such as 96-well plates or labelled beads.
Hybridization assays involve labelled nucleic acid probes to identify related DNA or RNA molecules (i.e. with significantly high degree of sequence similarity) within a complex mixture of unlabelled nucleic acid molecules. Antisense therapy, siRNA, and other oligonucleotide and nucleic acid based biotherapeutics can be quantified with hybridization assays.
Signalling of hybridization methods can be performed using oligonucleotide probes modified in-synthesis with haptens and small molecule ligands which act homologous to the capture and detection antibodies. As with traditional ELISA, conjugates to horse radish peroxidase (HRP) or alkaline phosphatase (AP) can be used as secondary antibodies.
Sandwich hybridization assay
[edit]
In the sandwich hybridization ELISA assay format, the antigen ligand and antibodies in ELISA are replaced with a nucleic acid analyte, complementary oligonucleotide capture and detection probes.[2]
Generally, in the case of nucleic acid hybridization, monovalent salt concentration and temperature are controlled for hybridization and wash stringency, contrary to a traditional ELISA, where the salt concentration will usually be fixed for the binding and wash steps (i.e. PBS or TBS). Thus, optimal salt concentration in hybridization assays varies dependent upon the length and base composition of the analyte, capture and detection probes.
Competitive hybridization assay
[edit]The competitive hybridization assay[3] is similar to a traditional competitive immunoassay. Like other hybridization assays, it relies on complementarity, where the capture probe competes between the analyte and the tracer–a labelled oligonucleotide analog to the analyte.
Hybridization-ligation assay
[edit]In the hybridization-ligation assay[4][5] a template probe replaces the capture probe in the sandwich assay for immobilization to the solid support. The template probe is fully complementary to the oligonucleotide analyte and is intended to serve as a substrate for T4 DNA ligase-mediated ligation. The template probe has in addition an additional stretch complementary to a ligation probe so that the ligation probe will ligate onto the 3'-end of the analyte. Albeit generic, the ligation probe is similar to a detection probe in that it is labelled with, for example, digoxigenin for downstream signalling. Stringent, low/no salt wash will remove un-ligated products.
The ligation of the analyte to the ligation probe makes the method specific for the 3'-end of the analyte, ligation by T4 DNA ligase being much less efficient over a bulge loop, which would happen for a 3' metabolite N-1 version of the analyte, for example. The specificity of the hybridization-ligation assay for ligation at the 3'-end is particularly relevant because the predominant nucleases in blood are 3' to 5' exonucleases.
One limitation of the method is that it requires a free 3'-end hydroxyl which may not be available when targeting moieties are attached to the 3'-end, for example. Further, more exotic nucleic acid chemistries with oligonucleotide drugs may impact upon the activity of the ligase, which needs to be determined on a case-by-case basis.
Dual ligation hybridization assay
[edit]
The dual ligation hybridization assay (DLA)[6] extends the specificity of the hybridization-ligation assay to a specific method for the parent compound. Despite hybridization-ligation assay's robustness, sensitivity and added specificity for the 3'-end of the oligonculeotide analyte, the hybridization-ligation assay is not specific for the 5' end of the analyte.
The DLA is intended to quantify the full-length, parent oligonucleotide compound only, with both intact 5' and 3' ends. DLA probes are ligated at the 5' and 3' ends of the analyte by the joint action of T4 DNA ligase and T4 polynucleotide kinase. The kinase phosphorylates the 5'-end of the analyte and the ligase will join the capture probe to the analyte to the detection probe. The capture and detection probes in the DLA can thus be termed ligation probes. As for the hybridization-ligation assay, the DLA is specific for the parent compound because the efficiency of ligation over a bulge loop is low, and thus the DLA detects the full-length analyte with both intact 5' and 3'-ends. The DLA can also be used for the determination of individual metabolites in biological matrices.
The limitations with the hybridization-ligation assay also apply to the dual ligation assay, with the 5'-end in addition to the 3'-end requiring to have a free hydroxyl (or a phosphate group). Further, T4 DNA ligases are incompatible with ligation of RNA molecules as a donor (i.e. RNA at the 5' end of the ligation). Therefore, second generation antisense that comprise 2'-O-methyl RNA, 2'-O-methoxyethyl or locked nucleic acids may not be compatible with the dual ligation assay.
Nuclease hybridization assay
[edit]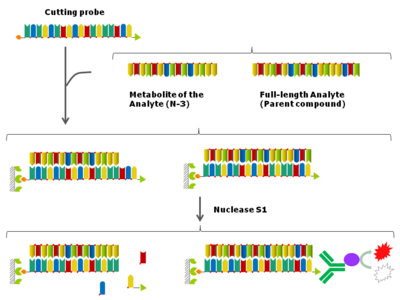
The nuclease hybridization assay,[7][8] also called S1 nuclease cutting assay, is a nuclease protection assay-based hybridization ELISA. The assay is using S1 nuclease, which degrades single-stranded DNA and RNA into oligo- or mononucleotides, leaving intact double-stranded DNA and RNA.
In the nuclease hybridization assay, the oligonucleotide analyte is captured onto the solid support such as a 96-well plate via a fully complementary cutting probe. After enzymatic processing by S1 nuclease, the free cutting probe and the cutting probe hybridized to metabolites, i.e. shortmers of the analyte are degraded, allowing signal to be generated only from the full-length cutting probe-analyte duplex.
The assay is well tolerant to diverse chemistries, as exemplified by the development of a nuclease assay for morpholino oligonucleotides.[9]
This assay set-up can lack robustness and is not suitable for validation following the FDA's guidelines for bioanalytical method validation. This is demonstrated by an absence of published method that have been validated to the standards outlined by the FDA for bioanalytical methods.
References
[edit]- ^ kim, Kalitsis, ji hun, paul; et al. "Nucleic Acids: Hybridisation". wiley. Retrieved 4 February 2017.
{{cite web}}: CS1 maint: multiple names: authors list (link) - ^ Efler, S.M.; Zhang, L.; Noll, B.O.; Uhlmann, E.; Davis, H.L. (2005), "Quantification of oligodeoxynucleotides in human plasma with a novel hybridization assay offers greatly enhanced sensitivity over capillary gel electrophoresis", Oligonucleotides, 15 (2): 119–131, doi:10.1089/oli.2005.15.119, PMID 15989426
- ^ Deverre, Jean-Robert; Boutet, Valérie; Boquet, Didier; Ezan, Eric; Grassi, Jacques; Grognet, Jean-Marc (1997), "A competitive enzyme hybridization assay for plasma determination of phosphodiester and phosphorothioate antisense oligonucleotides", Nucleic Acids Research, 25 (18): 3584–3589, doi:10.1093/nar/25.18.3584, PMC 146941, PMID 9278477[dead link]
- ^ Deverre, Jean-Robert; Boutet, Valérie; Boquet, Didier; Ezan, Eric; Grassi, Jacques; Grognet, Jean-Marc (1997), "A competitive enzyme hybridization assay for plasma determination of phosphodiester and phosphorothioate antisense oligonucleotides", Nucleic Acids Research, 25 (18): 3584–3589, doi:10.1093/nar/25.18.3584, PMC 146941, PMID 9278477[dead link]
- ^ US patent 6355438, Brenda F. Baker; Zhengrong Yu & Janet M. Leeds, "Method for quantitating oligonucleotides", published 2002-03-12, assigned to Isis Pharmaceuticals, Inc
- ^ Tremblay, Guy A.; Khalafaghian, Garinée; Legault, Julie; Bartlett, Alan (March 2011), "Dual ligation hybridization assay for the specific determination of oligonucleotide therapeutics", Bioanalysis, 3 (5): 499–508, doi:10.4155/bio.11.18, PMID 21388263
- ^ US patent 8163477, Zhengrong Yu; Brenda F. Baker & Hongjiang Wu, "Nuclease-based method for detecting and quantitating oligonucleotides", published 2012-04-24, assigned to Isis Pharmaceuticals, Inc
- ^ Xiaohui, Wei; Dai, Guowei; Marcucci, Guido; Liu, Zhongfa; Hoyt, Dale; Blum, William; Chan, Kenneth K. (2006), "A Specific Picomolar Hybridization-Based ELISA Assay for the Determination of Phosphorothioate Oligonucleotides in Plasma and Cellular Matrices", Pharmaceutical Research, 23 (6): 1251–1264, doi:10.1007/s11095-006-0082-3, PMID 16718617, S2CID 37490274
- ^ Burki, Umar; Keane, Jonathan; Blain, Alison; O'Donovan, Liz; Gait, Michael J.; Laval, Steven H.; Straub, Volker (2015), "Development and Application of an Ultrasensitive Hybridization-Based ELISA Method for the Determination of Peptide-Conjugated Phosphorodiamidate Morpholino Oligonucleotides", Nucleic Acid Therapeutics, 25 (5): 275–284, doi:10.1089/nat.2014.0528, PMC 4576940, PMID 26176274
