Heme oxygenase
| heme oxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
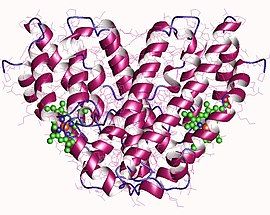 Heme oxygenase I, homodimer, Human | |||||||||
| Identifiers | |||||||||
| EC no. | 1.14.99.3 | ||||||||
| CAS no. | 9059-22-7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Heme oxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
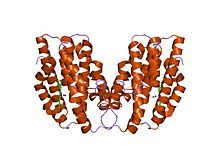 Crystal structures of ferrous and ferrous-no forms of verdoheme in a complex with human heme oxygenase-1: catalytic implications for heme cleavage | |||||||||
| Identifiers | |||||||||
| Symbol | Heme_oxygenase | ||||||||
| Pfam | PF01126 | ||||||||
| Pfam clan | CL0230 | ||||||||
| InterPro | IPR016053 | ||||||||
| PROSITE | PDOC00512 | ||||||||
| SCOP2 | 1qq8 / SCOPe / SUPFAM | ||||||||
| Membranome | 532 | ||||||||
| |||||||||
Heme oxygenase, or haem oxygenase, (HMOX, commonly abbreviated as HO) is an enzyme that catalyzes the degradation of heme to produce biliverdin, ferrous iron, and carbon monoxide.[1]
There are many heme degrading enzymes in nature. In general, only aerobic heme degrading enzymes are referred to as HMOX-like enzymes whereas anaerobic enzymes are typically not affiliated with the HMOX family.
Heme oxygenase
[edit]Heme oxygenase (alternatively spelled using haem or oxidase) catalyzes the degradation of heme to biliverdin/bilirubin, ferrous ion, and carbon monoxide. The human genome may encode three isoforms of HMOX.
The degradation of heme forms three distinct chromogens as seen in healing cycle of a bruise. This reaction can occur in virtually every cell and platelet; the classic example is the healing process of a contusion, which forms different chromogens as it gradually heals: (red) heme to (green) biliverdin to (yellow) bilirubin which is widely known for jaundice.[2] In general, aside from sharing the functionality of catabolizing heme, all HMOX isoforms share are signature 24-residue sequence considered to be essential for the enzymatic activity.[3]
Though present throughout the body, HMOX is most active in the spleen facilitating degradation of hemoglobin during erythrocyte recycling (approximately 0.8% of the erythrocyte pool per day).[4]
Heme oxygenase 1
[edit]Heme oxygenase 1 (HMOX1, commonly HO-1) is a member of the heat shock protein (HSP) family identified as HSP32. HO-1 is a 32kDa enzyme which contains 288 amino acid residues encoded by the HMOX1 gene. HO-1 is not a hemoprotein as it does not contain any heme prosthetic groups.[5] The activity of HO-1 is dependent upon NADPH-Cytochrome P450 Reductase.[6]
HO-1 is a stress-induced isoform present throughout the body[7] with highest concentrations in the spleen, liver, and kidneys, and on the cellular level is primarily located in the endoplasmic reticulum, although it has also been reported in the mitochondria, cell nucleus, and plasma membrane.[8] Soluble variations of HO-1 have been described. HO-1 may also serve as a chaperone protein, engage in protein-protein interactions, be secreted into the extracellular space, and participate in other cellular functions beyond its catalytic activity.[9] HO-1 may also generate small amounts of carbon suboxide.[10] HO-1 enzymes are degraded via ubiquitination.
The enzyme has been the subject of extensive investigation into its regulatory signaling, immunomodulatory, and cryoprotective roles.[11] HMOX1 is an essential enzyme. Human HMOX1-deficiency is rare, however several cases have been reported which generally results in death.[12]
In certain diseases, HMOX is problematic.[13][14] For example, HMOX1 may counteract certain chemotherapeutic drugs to rescue cancer cells from cytotoxic drugs thereby enabling cancer progression.[15] HMOX1 inhibitors are in development.[16]
Heme oxygenase 2
[edit]Heme oxygenase 2 (HMOX2 or HO-2) is a constitutive isoform that is expressed under homeostatic conditions in the testes, gastrointestinal tract, endothelial cells and the brain.[17] HO-2 is encoded by the HMOX2 gene. HO-2 is 36 kDa and shares 47% similarity with the HO-1 amino acid sequence; notably HO-2 has an extra N-terminal stretch of 20 amino acid residues.[5] Unlike HO-1, HO-2 is a hemoprotein containing heme regulatory motifs that contain heme independent of the heme catabolic site.[3]
Whereas HO-1 has innumerable inducers, only adrenal glucocorticoids are known to induce HO-2[12] whereas certain other molecules may increase its catalytic velocity.[9] Opioids may inhibit HMOX2 activity.[9] Many drugs that activate and inhibit HO-2 are in development.[18]
Heme oxygenase 3
[edit]A controversial third heme oxygenase (HO-3) is considered to be catalytically inactive and has been speculated to work in heme sensing or detoxification. HO-3 is 33 kDa with greatest presence in the liver, prostate, and kidneys. However, attempts to isolate HO-3 yielded pseudogenes derived from HO-2 transcripts thereby raising questions about the existence of a third isoform.[9]
Microbial heme oxygenase
[edit]Heme oxygenase is conserved across phylogenetic kingdoms.[19] The human microbiome contains dozens of unique microbial HMOX homologues which use many different abbreviations exemplified by:[9]
- HMX1 in Saccharomyces cerevisiae
- HmuO in Corynebacterium diphtheriae
- ChuS in commensal strains of Escherichia coli
A critical role of the prokaryotic HMOX systems is to facilitate acquisition of nutritional iron from a eukaryotic host.[20]
Some heme-degrading prokaryotic enzymes produce products such as formaldehyde rather than CO. As an example, certain pathogens such as Escherichia coli O157:H7 can express the non-CO producing ChuW isoform. Many pathogens are susceptible to CO toxicity, therefore expressing non-CO producing heme degradation enzymes evades self-inflicted toxicity while meeting nutritional iron needs. Commensal microbiota generally have CO tolerance as they produce and respond to CO signals; upon excretion from the microbe, the CO either directly benefits the host or applies selection pressure against pathogens thereby serving as a symbiotic currency.[9]
Plant heme oxygenase
[edit]Plants contain HMOX homologues with critical roles in plant physiology.[21] Although chlorophyll is structurally similar to heme, it is unclear if any HMOX-like enzymes are capable of facilitating metabolism.[9]
Quasi-enzymatic heme oxidation
[edit]As heme oxygenase is an enzymatic catalyst that accelerates the slow natural oxidation of heme, non-enzymatic oxidative degradation of heme, commonly termed 'coupled oxidation', was proposed as early as 1949. Akin to HMOX, coupled oxidation occurs at the alpha-methine bridge and leads to formation of biliverdin although the reaction's stoichiometry is different.[22] The first attempt to describe HMOX in 1962 by Nakajima turned out to be a non-enzymatic pathway. The complexity of the non-enzymatic pathway has been coined quasi-enzymatic or pseudoenzymatic.[22] A variety of mechanisms have been proposed.[22][23]
Reaction
[edit]HMOX1 is the rate-limiting step of heme catabolism that is dependent on NADPH-cytochrome P450 reductase and oxygen to cleave heme/porphyrin ring at the alpha-methene bridge to form biliverdin (or verdoglobin if the heme is still intact as hemoglobin). The reaction comprises three steps, which may be:[24]
- Heme b3+ + O
2 + NADPH + H+
→ α-meso-hydroxyheme3+ + NADP+
+ H
2O - α-meso-hydroxyheme3+ + H+
+ O
2 → verdoheme4+ + CO + H
2O - verdoheme4+ + 7/2 NADPH + O
2+ 3/2 H+
→ biliverdin + Fe2+ + 7/2 NADP+
+ H
2O
- Heme b3+ + O
The sum of these reactions is:
- Heme b3+ + 3O
2 + 9/2 NADPH + 7/2 H+
→ biliverdin + Fe2+ + CO + 9/2 NADP+
+ 3H
2O
If the iron is initially in the +2 state, the reaction could be:
- Heme b2+ + 3O2 + 4 NADPH + 4 H+ → biliverdin + Fe2+ + CO + 4 NADP+ + 3H2O
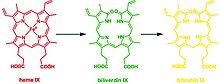
This reaction can occur in virtually every cell; the classic example is the formation of a contusion, which forms different chromogens as it gradually heals: (red) heme to (green) biliverdin to (yellow) bilirubin. In terms of molecular mechanisms, the enzyme facilitates the intramolecular hydroxylation of one meso carbon centre in the heme.[25]
Modulators
[edit]Inducers
[edit]HMOX1 is induced by countless molecules including heavy metals, statins, paclitaxel, rapamycin, probucol, nitric oxide, sildenafil, carbon monoxide, carbon monoxide-releasing molecules, and certain porphyrins.[26]
Phytochemical inducers of HO include: curcumin, resveratrol, piceatannol, caffeic acid phenethyl ester, dimethyl fumarate, fumaric acid esters, flavonoids, chalcones, ginkgo biloba, anthrocyanins, phlorotannins, carnosol, rosolic acid, and numerous other natural products.[26][27]
Endogenous inducers include i) lipids such as lipoxin and epoxyeicosatrienoic acid; and ii) peptides such as adrenomedullin and apolipoprotein; and iii) hemin.[26]
NRF2 inducers with downstream HO-1 induction include: genistein, 3-hydroxycoumarin, oleanolic acid, isoliquiritigenin, PEITC, diallyl trisulfide, oltipraz, benfotiamine, auranofin, acetaminophen, nimesulide, paraquat, ethoxyquin, diesel exhaust particles, silica, nanotubes, 15-deoxy-Δ12,14 prostaglandin J2, nitro-oleic acid, hydrogen peroxide, and succinylacetone.[28]
Inhibitors
[edit]HMOX1 is inhibited by certain porphyrins such as zinc protoporphyrin.[29]
Roles in physiology
[edit]HMOX is involved in numerous cellular operations.[30][31] The cyto-protective benefits of HMOX has stimulated significant research into its therapeutic and pharmaceutical potential.[32] These effects have not been verified in clinical trials.[33][8]
Carbon monoxide
[edit]HMOX is the main source of endogenous CO production,[33] though other minor contributors have been identified in recent years. CO is formed at a rate of 16.4 μmol/hour in the human body, ~86% originating from heme via heme oxygenase and ~14% from non-heme sources including: photooxidation, lipid and keto acid peroxidation, microbiome, and xenobiotics.[9] The average carboxyhemoglobin (CO-Hb) level in a non-smoker is between 0.2% and 0.85% CO-Hb (whereas a smoker may have between 4% and 10% CO-Hb), though genetics, geographic location, occupation, health and behavior are contributing variables.
Erythrocyte recycling in the spleen accounts for ~80% of heme-derived endogenous CO production. The remaining 20% of heme-derived CO production is attributed to hepatic catabolism of hemoproteins (myoglobin, cytochromes, catalase, peroxidases, soluble guanylate cyclase, nitric oxide synthase) and ineffective erythropoiesis in bone marrow.[4]
In addition to being a source of carbon monoxide, heme is a critical signal transducer involved in carbon monoxide sensing.[34][35] As a signaling agent, carbon monoxide is involved in normal physiology and has therapeutic benefits in many indications such as ameliorating inflammation and hypoxia.[33][36] It remain under investigation, however, to what extent HMOX is involved in carbon monoxide's protective effect against hypoxia as 3 molar equivalents of oxygen are required to produce carbon monoxide from heme catabolism, along with the question of heme bioavailability,[37] and slow HMOX1 induction which may take several hours (e.g. the slow healing of a bruise).[38]
Biliverdin / bilirubin
[edit]Ancient documentation for endogenous bilirubin traces back to medical humours written by Hippocrates.[39]
In most cases, HMOX selectively cleaves heme (iron protoporphyrin IX) at the α-methine bridge. The resulting bilirubin contains the suffix IXα to identify the composition of its structure by indicating its parent molecule was protoporphyrin IX cleaved at the alpha position (see protoporphyrin IX for further information on the Fischer nomenclature system). Drosophila melanogaster contains a unique HMOX that is not alpha specific resulting in formation of biliverdin IXα, IXβ, IXδ.[5] Non-enzymatic oxidation of heme is likewise non-specific resulting in ring opening at the α, β, γ, or δ positions.[22]
Biliverdin IXα undergoes biotransformation via biliverdin reductase to form bilirubin IXα.[2] Bilins play important roles across phylogenetic kingdoms.[40][41]
Ferrous ion
[edit]Ferrous ion is a common nomenclature used in the HMOX field for Iron(II) which appears in PubChem.[42] The iron liberated from HMOX is thought to be rapidly sequestered by ferritin. However, reactive oxygen species generated through the Fenton or Haber-Weiss reactions may enable downstream signaling.[43][44]
History
[edit]HMOX1 was first characterized by Tenhunen and Rudi Schmid upon demonstrating it as the enzyme responsible for catalyzing biotransformation of heme to bilirubin.[12]
Several labs attempted to explain the biotransformation of heme to biliverdin such as Nakajima et al. in 1962 who characterized a soluble "heme α-methenyl oxygenase", however the findings could not be reproduced and alternative non-enzymatic explanations for their observation emerged. The earliest evidence of oxidative enzymatic biotransformation of heme to a bilin was demonstrated by Hans Plieninger and Hans Fischer in 1942.[45] The discovery of HMOX is a unique case of academic lineage as Fischer was the academic adviser for Cecil Watson, and Watson was an adviser for Rudi Schmid.
Felix Hoppe-Seyler coined the name "haemoglobin"; haem being derived from Greek meaning blood, and globin from Latin globus meaning round object (see also: carboxyhemoglobin etymology). Hemoglobin was first discovered in the 1840s by Friedrich Ludwig Hünefeld.[46][47] Heme (as hemin coordinated with chlorine) was characterized by Ludwik Karol Teichmann in 1853. Many labs investigated in vitro transformation of heme into bilins throughout the 1930s exemplified by the work of Georg Barkan,[48] followed by Esther Killick who recognized a presence of carbon monoxide to correlate with pseudohemoglobin (an obsolete bilin term coined by Barkan) in 1940.[12] The endogenous biotransformation of heme to bilirubin is thought to have been definitively demonstrated with experimental evidence by Irving London in 1950,[49] although trace evidence for the endogenous formation of bilirubin has origins dating back several centuries in the context of jaundice with innumerable global contributions (see also: History of Bilirubin).[2][45]
CO was detected in exhaled breath 1869.[12] Felix Hoppe-Seyler developed the first qualitative carboxyhemoglobin test, and Josef von Fodor developed the first quantitative analytical test for carboxyhemoglobin.[12] The first reported detection of naturally occurring CO in human blood occurred in 1923 by Royd Ray Sayers et al. although they discarded their data as random error.[12] Alexander Gettler confirmed CO to have a normal presence in blood in 1933, however, he attributed the finding to inevitable pollution exposure or perhaps derived from the human microbiome.[9] Sjöstrand later demonstrated CO production from hemoglobin decomposition in 1952.[12]
References
[edit]- ^ Ryter SW, Alam J, Choi AM (April 2006). "Heme oxygenase-1/carbon monoxide: from basic science to therapeutic applications". Physiological Reviews. 86 (2): 583–650. doi:10.1152/physrev.00011.2005. PMID 16601269.
- ^ a b c Wegiel B, Otterbein LE (2012). "Go green: the anti-inflammatory effects of biliverdin reductase". Frontiers in Pharmacology. 3: 47. doi:10.3389/fphar.2012.00047. PMC 3306015. PMID 22438844.
- ^ a b McCoubrey WK, Huang TJ, Maines MD (May 1997). "Heme oxygenase-2 is a hemoprotein and binds heme through heme regulatory motifs that are not involved in heme catalysis". The Journal of Biological Chemistry. 272 (19): 12568–12574. doi:10.1074/jbc.272.19.12568. PMID 9139709.
- ^ a b Vreman H, Wong R, Stevenson D (2001-10-30). "Sources, Sinks, and Measurement of Carbon Monoxide". Carbon Monoxide and Cardiovascular Functions. CRC Press. pp. 273–307. doi:10.1201/9781420041019.ch15 (inactive 2024-11-11). ISBN 978-0-8493-1041-6.
{{cite book}}: CS1 maint: DOI inactive as of November 2024 (link) - ^ a b c Kikuchi G, Yoshida T, Noguchi M (December 2005). "Heme oxygenase and heme degradation". Biochemical and Biophysical Research Communications. 338 (1): 558–567. doi:10.1016/j.bbrc.2005.08.020. PMID 16115609.
- ^ Wang J, de Montellano PR (May 2003). "The binding sites on human heme oxygenase-1 for cytochrome p450 reductase and biliverdin reductase". The Journal of Biological Chemistry. 278 (22): 20069–20076. doi:10.1074/jbc.M300989200. PMID 12626517.
- ^ Otterbein LE, Choi AM (December 2000). "Heme oxygenase: colors of defense against cellular stress". American Journal of Physiology. Lung Cellular and Molecular Physiology. 279 (6): L1029-37. doi:10.1152/ajplung.2000.279.6.L1029. PMID 11076792. S2CID 8813119.
- ^ a b Hopper CP, Meinel L, Steiger C, Otterbein LE (2018-10-11). "Where is the Clinical Breakthrough of Heme Oxygenase-1 / Carbon Monoxide Therapeutics?". Current Pharmaceutical Design. 24 (20): 2264–2282. doi:10.2174/1381612824666180723161811. PMID 30039755. S2CID 51712930.
- ^ a b c d e f g h i Hopper CP, De La Cruz LK, Lyles KV, Wareham LK, Gilbert JA, Eichenbaum Z, et al. (December 2020). "Role of Carbon Monoxide in Host-Gut Microbiome Communication". Chemical Reviews. 120 (24): 13273–13311. doi:10.1021/acs.chemrev.0c00586. PMID 33089988. S2CID 224824871.
- ^ Tsoureas N, Green JC, Cloke FG, Puschmann H, Roe SM, Tizzard G (June 2018). "Trimerisation of carbon suboxide at a di-titanium centre to form a pyrone ring system". Chemical Science. 9 (22): 5008–5014. doi:10.1039/c8sc01127c. PMC 5994745. PMID 29938029.
- ^ Campbell NK, Fitzgerald HK, Dunne A (January 2021). "Regulation of inflammation by the antioxidant haem oxygenase 1". Nature Reviews. Immunology. 21 (7): 411–425. doi:10.1038/s41577-020-00491-x. PMID 33514947. S2CID 231762031.
- ^ a b c d e f g h Hopper CP, Zambrana PN, Goebel U, Wollborn J (June 2021). "A brief history of carbon monoxide and its therapeutic origins". Nitric Oxide. 111–112: 45–63. doi:10.1016/j.niox.2021.04.001. ISSN 1089-8603. PMID 33838343. S2CID 233205099.
- ^ Schipper HM, Song W, Tavitian A, Cressatti M (January 2019). "The sinister face of heme oxygenase-1 in brain aging and disease". Progress in Neurobiology. 172: 40–70. doi:10.1016/j.pneurobio.2018.06.008. PMID 30009872. S2CID 51629548.
- ^ Jozkowicz A, Was H, Dulak J (December 2007). "Heme oxygenase-1 in tumors: is it a false friend?". Antioxidants & Redox Signaling. 9 (12): 2099–117. doi:10.1089/ars.2007.1659. PMC 2096718. PMID 17822372.
- ^ Podkalicka P, Mucha O, Józkowicz A, Dulak J, Łoboda A (March 2018). "Heme oxygenase inhibition in cancers: possible tools and targets". Contemporary Oncology. 22 (1A): 23–32. doi:10.5114/wo.2018.73879. PMC 5885082. PMID 29628790.
- ^ Salerno L, Floresta G, Ciaffaglione V, Gentile D, Margani F, Turnaturi R, et al. (April 2019). "Progress in the development of selective heme oxygenase-1 inhibitors and their potential therapeutic application". European Journal of Medicinal Chemistry. 167: 439–453. doi:10.1016/j.ejmech.2019.02.027. PMID 30784878. S2CID 73496388.
- ^ Muñoz-Sánchez J, Chánez-Cárdenas ME (2014). "A review on hemeoxygenase-2: focus on cellular protection and oxygen response". Oxidative Medicine and Cellular Longevity. 2014: 604981. doi:10.1155/2014/604981. PMC 4127239. PMID 25136403.
- ^ Intagliata S, Salerno L, Ciaffaglione V, Leonardi C, Fallica AN, Carota G, et al. (December 2019). "Heme Oxygenase-2 (HO-2) as a therapeutic target: Activators and inhibitors". European Journal of Medicinal Chemistry. 183: 111703. doi:10.1016/j.ejmech.2019.111703. PMID 31550661. S2CID 202760666.
- ^ Li C, Stocker R (2009). "Heme oxygenase and iron: from bacteria to humans". Redox Report. 14 (3): 95–101. doi:10.1179/135100009X392584. PMID 19490750. S2CID 206840483.
- ^ Frankenberg-Dinkel N (October 2004). "Bacterial heme oxygenases". Antioxidants & Redox Signaling. 6 (5): 825–34. doi:10.1089/ars.2004.6.825. PMID 15345142.
- ^ He H, He L (2014). "Heme oxygenase 1 and abiotic stresses in plants". Acta Physiologiae Plantarum. 36 (3): 581–588. doi:10.1007/s11738-013-1444-1. ISSN 0137-5881. S2CID 6842488.
- ^ a b c d O'Carra P, Colleran E (1977). "Chapter 3: Nonenzymatic and quasi-enzymic models for catabolic heme cleavage". In Berk PD, Berlin NI (eds.). International Symposium on Chemistry and Physiology of Bile Pigments. U.S. Department of Health, Education, and Welfare, Public Health Service, National Institutes of Health. pp. 26–40.
- ^ Berk PD, Berlin NI (1977). "Chapter 4: Mechanism of the ring opening of heme". International Symposium on Chemistry and Physiology of Bile Pigments. U.S. Department of Health, Education, and Welfare, Public Health Service, National Institutes of Health. pp. 42–66.
- ^ Evans JP, Niemevz F, Buldain G, de Montellano PO (July 2008). "Isoporphyrin intermediate in heme oxygenase catalysis. Oxidation of alpha-meso-phenylheme". The Journal of Biological Chemistry. 283 (28): 19530–9. doi:10.1074/jbc.M709685200. PMC 2443647. PMID 18487208. The reference does not give the exact stoichiometry of each reaction.
- ^ Yoshida T, Migita CT (November 2000). "Mechanism of heme degradation by heme oxygenase". Journal of Inorganic Biochemistry. 82 (1–4): 33–41. doi:10.1016/S0162-0134(00)00156-2. PMID 11132636.
- ^ a b c Ferrándiz ML, Devesa I (2008). "Inducers of heme oxygenase-1". review. Current Pharmaceutical Design. 14 (5): 473–86. doi:10.2174/138161208783597399. PMID 18289074.
- ^ Correa-Costa M, Otterbein LE (2014). "Eat to Heal: Natural Inducers of the Heme Oxygenase-1 System". In Folkerts G, Garssen J (eds.). Pharma-Nutrition. secondary. AAPS Advances in the Pharmaceutical Sciences Series. Vol. 12. Springer, Cham. pp. 243–256. doi:10.1007/978-3-319-06151-1_12. ISBN 978-3-319-06150-4.
- ^ Ma Q, He X (October 2012). "Molecular basis of electrophilic and oxidative defense: promises and perils of Nrf2". Pharmacological Reviews. 64 (4): 1055–81. doi:10.1124/pr.110.004333. PMC 4648289. PMID 22966037.
- ^ Vreman HJ, Cipkala DA, Stevenson DK (March 1996). "Characterization of porphyrin heme oxygenase inhibitors". Canadian Journal of Physiology and Pharmacology. 74 (3): 278–85. PMID 8773407.
- ^ Kim HJ, Joe Y, Surh YJ, Chung HT (December 2018). "Metabolic signaling functions of the heme oxygenase/CO system in metabolic diseases". Cellular & Molecular Immunology. 15 (12): 1085–1087. doi:10.1038/s41423-018-0045-8. PMC 6269490. PMID 29807990.
- ^ Ryter SW, Otterbein LE, Morse D, Choi AM (2002-05-01). "Heme oxygenase/carbon monoxide signaling pathways: regulation and functional significance". Molecular and Cellular Biochemistry. 234–235 (1–2): 249–63. doi:10.1023/A:1015957026924. PMC 7101540. PMID 12162441.
- ^ Cheng Y, Rong J (2017-10-03). "Therapeutic Potential of Heme Oxygenase-1/carbon Monoxide System Against Ischemia-Reperfusion Injury". Current Pharmaceutical Design. 23 (26): 3884–3898. doi:10.2174/1381612823666170413122439. PMID 28412905.
- ^ a b c Motterlini R, Otterbein LE (September 2010). "The therapeutic potential of carbon monoxide". Nature Reviews. Drug Discovery. 9 (9): 728–43. doi:10.1038/nrd3228. PMID 20811383. S2CID 205477130.
- ^ Shimizu T, Lengalova A, Martínek V, Martínková M (December 2019). "Heme: emergent roles of heme in signal transduction, functional regulation and as catalytic centres". Chemical Society Reviews. 48 (24): 5624–5657. doi:10.1039/C9CS00268E. PMID 31748766. S2CID 208217502.
- ^ Shimizu T, Huang D, Yan F, Stranava M, Bartosova M, Fojtíková V, Martínková M (July 2015). "Gaseous O2, NO, and CO in signal transduction: structure and function relationships of heme-based gas sensors and heme-redox sensors". Chemical Reviews. 115 (13): 6491–533. doi:10.1021/acs.chemrev.5b00018. PMID 26021768.
- ^ Motterlini R, Foresti R (March 2017). "Biological signaling by carbon monoxide and carbon monoxide-releasing molecules". American Journal of Physiology. Cell Physiology. 312 (3): C302 – C313. doi:10.1152/ajpcell.00360.2016. PMID 28077358.
- ^ Leung GC, Fung SS, Gallio AE, Blore R, Alibhai D, Raven EL, Hudson AJ (June 2021). "Unravelling the mechanisms controlling heme supply and demand". Proceedings of the National Academy of Sciences of the United States of America. 118 (22): e2104008118. Bibcode:2021PNAS..11804008L. doi:10.1073/pnas.2104008118. PMC 8179208. PMID 34035176.
- ^ Pellacani A, Wiesel P, Sharma A, Foster LC, Huggins GS, Yet SF, Perrella MA (August 1998). "Induction of heme oxygenase-1 during endotoxemia is downregulated by transforming growth factor-beta1". Circulation Research. 83 (4): 396–403. doi:10.1161/01.RES.83.4.396. PMID 9721696.
- ^ Watson, Cecil J. (1977). "Historical Review of Bilirubin Chemistry". In Berk, Paul D. (ed.). International Symposium on Chemistry and Physiology of Bile Pigments. U.S. Department of Health, Education, and Welfare, Public Health Service, National Institutes of Health. pp. 3–16
- ^ Shekhawat GS (2019). eLS (1st ed.). Wiley. doi:10.1002/9780470015902.a0028352. ISBN 978-0-470-01617-6. S2CID 146029918.
- ^ Takemoto JY, Chang CW, Chen D, Hinton G (2019). "Heme-Derived Bilins". Israel Journal of Chemistry. 59 (5): 378–386. doi:10.1002/ijch.201800167. ISSN 0021-2148. S2CID 104394477.
- ^ "Ferrous ion". PubChem. Retrieved 2021-05-26.
- ^ Sies H, Jones DP (July 2020). "Reactive oxygen species (ROS) as pleiotropic physiological signalling agents". Nature Reviews. Molecular Cell Biology. 21 (7): 363–383. doi:10.1038/s41580-020-0230-3. PMID 32231263. S2CID 214695993.
- ^ NaveenKumar SK, SharathBabu BN, Hemshekhar M, Kemparaju K, Girish KS, Mugesh G (August 2018). "The Role of Reactive Oxygen Species and Ferroptosis in Heme-Mediated Activation of Human Platelets". ACS Chemical Biology. 13 (8): 1996–2002. doi:10.1021/acschembio.8b00458. PMID 29869870. S2CID 46936967.
- ^ a b Watson C (1977). "Historical Review of Bilirubin Chemistry". In Berk P (ed.). Chemistry and Physiology of Bile Pigments. p. 5.
- ^ Clegg B (2011). "Haemoglobin". Chemistry World. Retrieved 2021-05-26.
- ^ Boor AK (January 1930). "A Crystallographic Study of Pure Carbonmon-Oxide Hemoglobin". The Journal of General Physiology. 13 (3): 307–316. doi:10.1085/jgp.13.3.307. PMC 2141039. PMID 19872525.
- ^ Barkan G, Schales O (November 1938). "A Hæmoglobin from Bile Pigment". Nature. 142 (3601): 836–837. Bibcode:1938Natur.142..836B. doi:10.1038/142836b0. ISSN 1476-4687. S2CID 4073510.
- ^ "Bilirubin". American Chemical Society. Retrieved 2021-10-19.
External links
[edit]- Heme+Oxygenase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- EC 1.14.99.3
