Canalisation (genetics)
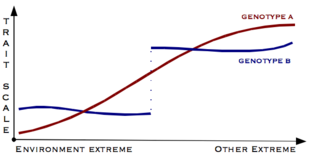
Canalisation is a measure of the ability of a population to produce the same phenotype regardless of variability of its environment or genotype. It is a form of evolutionary robustness. The term was coined in 1942 by C. H. Waddington to capture the fact that "developmental reactions, as they occur in organisms submitted to natural selection...are adjusted so as to bring about one definite end-result regardless of minor variations in conditions during the course of the reaction".[1] He used this word rather than robustness to consider that biological systems are not robust in quite the same way as, for example, engineered systems.
Biological robustness or canalisation comes about when developmental pathways are shaped by evolution. Waddington introduced the concept of the epigenetic landscape, in which the state of an organism rolls "downhill" during development. In this metaphor, a canalised trait is illustrated as a valley (which he called a creode) enclosed by high ridges, safely guiding the phenotype to its "fate". Waddington claimed that canals form in the epigenetic landscape during evolution, and that this heuristic is useful for understanding the unique qualities of biological robustness.[2]
Genetic assimilation
[edit]Waddington used the concept of canalisation to explain his experiments on genetic assimilation.[3] In these experiments, he exposed Drosophila pupae to heat shock. This environmental disturbance caused some flies to develop a crossveinless phenotype. He then selected for crossveinless. Eventually, the crossveinless phenotype appeared even without heat shock. Through this process of genetic assimilation, an environmentally induced phenotype had become inherited. Waddington explained this as the formation of a new canal in the epigenetic landscape.
It is, however, possible to explain genetic assimilation using only quantitative genetics and a threshold model, with no reference to the concept of canalisation.[4][5][6][7] However, theoretical models that incorporate a complex genotype–phenotype map have found evidence for the evolution of phenotypic robustness[8] contributing to genetic assimilation,[9] even when selection is only for developmental stability and not for a particular phenotype, and so the quantitative genetics models do not apply. These studies suggest that the canalisation heuristic may still be useful, beyond the more simple concept of robustness.
Congruence hypothesis
[edit]Neither canalisation nor robustness are simple quantities to quantify: it is always necessary to specify which trait is canalised (robust) to which perturbations. For example, perturbations can come either from the environment or from mutations. It has been suggested that different perturbations have congruent effects on development taking place on an epigenetic landscape.[10][11][12][13][14] This could, however, depend on the molecular mechanism responsible for robustness, and be different in different cases.[15]
Evolutionary capacitance
[edit]The canalisation metaphor suggests that some phenotypic traits are very robust to small perturbations, for which development does not exit the canal, and rapidly returns down, with little effect on the final outcome of development. But perturbations whose magnitude exceeds a certain threshold will break out of the canal, moving the developmental process into uncharted territory. For instance, the study of an allelic series for Fgf8, an important gene for craniofacial development, with decreasing levels of gene expression demonstrated that the phenotype remains canalised as long as the expression level is above 40% of the wild-type expression.[16]
Strong robustness up to a limit, with little robustness beyond, is a pattern that could increase evolvability in a fluctuating environment.[17] Canalisation of a large set of genotypes into a limited phenotypic space has been suggested as a mechanism for the accumulation, in a neutral manner, of mutations that could otherwise be deleterious.[18] Genetic canalisation could allow for evolutionary capacitance, where genetic diversity accumulates in a population over time, sheltered from natural selection because it does not normally affect phenotypes. This hidden diversity could then be unleashed by extreme changes in the environment or by molecular switches, releasing previously cryptic genetic variation that can then contribute to a rapid burst of evolution,[18] a phenomenon termed decanalisation. Cycles of canalization-decanalization could explain the alternating periods of stasis, where genotypic diversity accumulates without morphological changes, followed by rapid morphological changes, where decanalization releases the phenotypic diversity and becomes subject to natural selection, in the fossil record, thus providing a potential developmental explanation for the punctuated equilibrium.[17]
HSP90 and decanalisation
[edit]In 1998, Susan Lindquist discovered that Drosophila hsp83 heterozygous mutants exhibit a large diversity of phenotypes (from sexual combs on the head, to scutoid-like and notched wings phenotypes). She showed that these phenotypes could be passed on to the next generation, suggesting a genetic basis for those phenotypes.[19] The authors hypothesized that Hsp90 (the gene mutated in hsp83), as a chaperone protein, plays a pivotal role in the folding and activation of many proteins involved in developmental signaling pathways, thus buffering against genetic variation in those pathways.[20] hsp83 mutants would therefore release the cryptic genetic variation, resulting in a diversity of phenotypes.
In 2002, Lindquist showed that pharmacological inhibition of HSP90 in Arabidopsis thaliana also lead to a wide range of phenotypes, some of which could be considered adaptive, further supporting the canalising role of HSP90.[21]
Finally, the same type of experiment in the cavefish Astyanax mexicanus yielded similar results. This species encompasses two populations: an eyed population living under the water surface and an eye-less blind population living in caves. Not only is the cave population eye-less but it also displays a largely reduced orbit size. HSP90 inhibition leads to an increased variation in orbit size that could explain how this trait could evolve in just a few generations. Further analysis showed that low conductivity in the cave water induces a stress response mimicking the inhibition of HSP90, providing a mechanism for decanalisation.[22]
Interpretation of the original Drosophila paper[19] is now subject to controversy. Molecular analysis of the hsp83 mutant showed that HSP90 is required for piRNA biogenesis, a set of small RNAs repressing transposons in the germline.,[23] causing massive transposon[24] insertional mutagenesis that could explain the phenotypic diversification.[25]
Significance of Variability in Components
[edit]Understanding variability is an important aspect of comprehending natural selection and mutations. Variability can be classified into two categories: modulating phenotypic variation and modulating the phenotypes that are produced.[26] The presence of this so-called bias in genetic variability allows us to gain further insights into how certain phenotypes are more successful in terms of their actual morphology, biochemical makeup, or behavior.[27] It is scientifically known that organisms need to develop systematically integrated systems in order to thrive in their specific ecosystems. This extends to morphology, where variations must occur in a systematic order; otherwise, phenotypic mutations will not persist due to the occurrence of natural selection. The variation affects the speed and rate of evolutionary change through the selection and modulation of phenotypic variations.[28] Ultimately, this results in a lower amount of diversity observed throughout evolution, as the majority of phenotypes do not persist beyond a few generations due to their inferior morphology, biochemical makeup, or physical movement or appearance.
See also
[edit]- Developmental noise
- Phenotypic integration
- Phenotypic plasticity
- Developmental systems theory
- Gene regulatory network
- Systems biology
References
[edit]- ^ Waddington CH (1942). "Canalization of development and the inheritance of acquired characters". Nature. 150 (3811): 563–565. Bibcode:1942Natur.150..563W. doi:10.1038/150563a0. S2CID 4127926.
- ^ Waddington CH (1957). The strategy of the genes. George Allen & Unwin.
- ^ Waddington CH (1953). "Genetic assimilation of an acquired character". Evolution. 7 (2): 118–126. doi:10.2307/2405747. JSTOR 2405747.
- ^ Stern C (1958). "Selection for subthreshold differences and the origin of pseudoexogenous adaptations". American Naturalist. 92 (866): 313–316. doi:10.1086/282040. S2CID 84634317.
- ^ Bateman KG (1959). "The genetic assimilation of the dumpy phenocopy". American Naturalist. 56 (3): 341–351. doi:10.1007/bf02984790. S2CID 41242659.
- ^ Scharloo W (1991). "Canalization – genetic and developmental aspects". Annual Review of Ecology and Systematics. 22: 65–93. doi:10.1146/annurev.es.22.110191.000433.
- ^ Falconer DS, Mackay TF (1996). Introduction to Quantitative Genetics. pp. 309–310.
- ^ Siegal ML, Bergman A (August 2002). "Waddington's canalization revisited: developmental stability and evolution". Proceedings of the National Academy of Sciences of the United States of America. 99 (16): 10528–32. Bibcode:2002PNAS...9910528S. doi:10.1073/pnas.102303999. PMC 124963. PMID 12082173.
- ^ Masel J (September 2004). "Genetic assimilation can occur in the absence of selection for the assimilating phenotype, suggesting a role for the canalization heuristic". Journal of Evolutionary Biology. 17 (5): 1106–10. doi:10.1111/j.1420-9101.2004.00739.x. PMID 15312082.
- ^ Meiklejohn CD, Hartl DL (2002). "A single mode of canalization". Trends in Ecology & Evolution. 17 (10): 468–473. doi:10.1016/S0169-5347(02)02596-X.
- ^ Ancel LW, Fontana W (October 2000). "Plasticity, evolvability, and modularity in RNA". The Journal of Experimental Zoology. 288 (3): 242–83. CiteSeerX 10.1.1.43.6910. doi:10.1002/1097-010X(20001015)288:3<242::AID-JEZ5>3.0.CO;2-O. PMID 11069142.
- ^ Szöllosi GJ, Derényi I (April 2009). "Congruent evolution of genetic and environmental robustness in micro-RNA". Molecular Biology and Evolution. 26 (4): 867–74. arXiv:0810.2658. doi:10.1093/molbev/msp008. PMID 19168567.
- ^ Wagner GP, Booth G, Bagheri-Chaichian H (April 1997). "A Population Genetic Theory of Canalization". Evolution; International Journal of Organic Evolution. 51 (2): 329–347. CiteSeerX 10.1.1.27.1001. doi:10.2307/2411105. JSTOR 2411105. PMID 28565347.
- ^ Lehner B (February 2010). Polymenis M (ed.). "Genes confer similar robustness to environmental, stochastic, and genetic perturbations in yeast". PLOS ONE. 5 (2): e9035. Bibcode:2010PLoSO...5.9035L. doi:10.1371/journal.pone.0009035. PMC 2815791. PMID 20140261.
- ^ Masel J, Siegal ML (September 2009). "Robustness: mechanisms and consequences". Trends in Genetics. 25 (9): 395–403. doi:10.1016/j.tig.2009.07.005. PMC 2770586. PMID 19717203.
- ^ Green RM, Fish JL, Young NM, Smith FJ, Roberts B, Dolan K, et al. (December 2017). "Developmental nonlinearity drives phenotypic robustness". Nature Communications. 8 (1): 1970. Bibcode:2017NatCo...8.1970G. doi:10.1038/s41467-017-02037-7. PMC 5719035. PMID 29213092.
- ^ a b Eshel I, Matessi C (August 1998). "Canalization, genetic assimilation and preadaptation. A quantitative genetic model". Genetics. 149 (4): 2119–33. doi:10.1093/genetics/149.4.2119. PMC 1460279. PMID 9691063.
- ^ a b Paaby AB, Rockman MV (April 2014). "Cryptic genetic variation: evolution's hidden substrate". Nature Reviews. Genetics. 15 (4): 247–58. doi:10.1038/nrg3688. PMC 4737706. PMID 24614309.
- ^ a b Rutherford SL, Lindquist S (November 1998). "Hsp90 as a capacitor for morphological evolution". Nature. 396 (6709): 336–42. Bibcode:1998Natur.396..336R. doi:10.1038/24550. PMID 9845070. S2CID 204996106.
- ^ Whitesell L, Lindquist SL (October 2005). "HSP90 and the chaperoning of cancer". Nature Reviews. Cancer. 5 (10): 761–72. doi:10.1038/nrc1716. PMID 16175177. S2CID 22098282.
- ^ Queitsch C, Sangster TA, Lindquist S (June 2002). "Hsp90 as a capacitor of phenotypic variation". Nature. 417 (6889): 618–24. Bibcode:2002Natur.417..618Q. doi:10.1038/nature749. PMID 12050657. S2CID 4419085.
- ^ Rohner N, Jarosz DF, Kowalko JE, Yoshizawa M, Jeffery WR, Borowsky RL, et al. (December 2013). "Cryptic variation in morphological evolution: HSP90 as a capacitor for loss of eyes in cavefish". Science. 342 (6164): 1372–5. Bibcode:2013Sci...342.1372R. doi:10.1126/science.1240276. PMC 4004346. PMID 24337296.
- ^ Mutation in this gene results in the gene being expressed
- ^ Hackett, Perry B.; Largaespada, David A.; Switzer, Kirsten C.; Cooper, Laurence J. N. (1 April 2013). "Evaluating risks of insertional mutagenesis by DNA transposons in gene therapy". Translational Research. 161 (4): 265–283. doi:10.1016/j.trsl.2012.12.005. PMC 3602164. PMID 23313630.
- ^ Specchia V, Piacentini L, Tritto P, Fanti L, D'Alessandro R, Palumbo G, et al. (February 2010). "Hsp90 prevents phenotypic variation by suppressing the mutagenic activity of transposons". Nature. 463 (7281): 662–5. Bibcode:2010Natur.463..662S. doi:10.1038/nature08739. PMID 20062045. S2CID 4429205.
- ^ Hallgrímsson, Benedikt; Willmore, Katherine; Hall, Brian K. (2002). "Canalization, Developmental Stability, and Morphological Integration in Primate Limbs". American Journal of Physical Anthropology. Suppl 35: 131–158. doi:10.1002/ajpa.10182. PMC 5217179. PMID 12653311.
- ^ West-Eberhard, M J (November 2018). "Phenotypic Plasticity and the Origins of Diversity". Annual Review of Ecology and Systematics. 20 (Annual Review of Ecology and Systematics): 249–278. doi:10.1146/annurev.es.20.110189.001341.
- ^ Loewe, Laurence; Hill, William G. (27 April 2010). "The population genetics of mutations: good, bad and indifferent". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 365 (1544): 1153–1167. doi:10.1098/rstb.2009.0317. PMC 2871823. PMID 20308090.
