Baltimore classification
Baltimore classification is a system used to classify viruses based on their manner of messenger RNA (mRNA) synthesis. By organizing viruses based on their manner of mRNA production, it is possible to study viruses that behave similarly as a distinct group. Seven Baltimore groups are described that take into consideration whether the viral genome is made of deoxyribonucleic acid (DNA) or ribonucleic acid (RNA), whether the genome is single- or double-stranded, and whether the sense of a single-stranded RNA genome is positive or negative.
Baltimore classification also closely corresponds to the manner of replicating the genome, so Baltimore classification is useful for grouping viruses together for both transcription and replication. Certain subjects pertaining to viruses are associated with multiple, specific Baltimore groups, such as specific forms of translation of mRNA and the host range of different types of viruses. Structural characteristics such as the shape of the viral capsid, which stores the viral genome, and the evolutionary history of viruses are not necessarily related to Baltimore groups.
Baltimore classification was created in 1971 by virologist David Baltimore. Since then, it has become common among virologists to use Baltimore classification alongside standard virus taxonomy, which is based on evolutionary history. In 2018 and 2019, Baltimore classification was partially integrated into virus taxonomy based on evidence that certain groups were descended from common ancestors. Various realms, kingdoms, and phyla now correspond to specific Baltimore groups.
Overview
[edit]Baltimore classification groups viruses together based on their manner of mRNA synthesis. Characteristics directly related to this include whether the genome is made of deoxyribonucleic acid (DNA) or ribonucleic acid (RNA), the strandedness of the genome, which can be either single- or double-stranded, and the sense of a single-stranded genome, which is either positive or negative. The primary advantage of Baltimore classification is that by classifying viruses according to the aforementioned characteristics, viruses that behave in the same manner can be studied as distinct groups. There are seven Baltimore groups numbered with Roman numerals, listed hereafter.[1]
- Group I: double-stranded DNA viruses
- Group II: single-stranded DNA viruses
- Group III: double-stranded RNA viruses
- Group IV: positive sense single-stranded RNA viruses
- Group V: negative sense single-stranded RNA viruses
- Group VI: single-stranded RNA viruses with a DNA intermediate in their life cycle
- Group VII: double-stranded DNA viruses with an RNA intermediate in their life cycle
Baltimore classification is chiefly based on the transcription of the viral genome, and viruses within each group typically share the manners by which the mRNA synthesis occurs. While not the direct focus of Baltimore classification, groups are organized in such a manner that viruses in each group also typically have the same mechanisms of replicating the viral genome.[2][3] Because of this, Baltimore classification provides insights into both the transcription and replication parts of the viral life cycle. Structural characteristics of a virus particle, called a virion, such as the shape of the viral capsid and the presence of a viral envelope, a lipid membrane that surrounds the capsid, have no direct relation to Baltimore groups, nor do the groups necessarily show genetic relation based on evolutionary history.[1]

Classification
[edit]DNA viruses
[edit]DNA viruses have genomes made of deoxyribonucleic acid (DNA) and are organized into two groups: double-stranded DNA (dsDNA) viruses, and single-stranded DNA (ssDNA) viruses. They are assigned to four separate realms: Adnaviria, Duplodnaviria, Monodnaviria, and Varidnaviria. Many have yet to be assigned to a realm.
Group I: double-stranded DNA viruses
[edit]The first Baltimore group contains viruses that have a double-stranded DNA (dsDNA) genome. All dsDNA viruses have their mRNA synthesized in a three-step process. First, a transcription preinitiation complex binds to the DNA upstream of the site where transcription begins, allowing for the recruitment of a host RNA polymerase. Second, once the RNA polymerase is recruited, it uses the negative strand as a template for synthesizing mRNA strands. Third, the RNA polymerase terminates transcription upon reaching a specific signal, such as a polyadenylation site.[4][5][6]
dsDNA viruses make use of several mechanisms to replicate their genome. Bidirectional replication, in which two replication forks are established at a replication origin site and move in opposite directions of each other, is widely used.[7] A rolling circle mechanism that produces linear strands while progressing in a loop around the circular genome is also common.[8] Some dsDNA viruses use a strand displacement method whereby one strand is synthesized from a template strand, and a complementary strand is then synthesized from the prior synthesized strand, forming a dsDNA genome.[9] Lastly, some dsDNA viruses are replicated as part of a process called replicative transposition whereby a viral genome in a host cell's DNA is replicated to another part of a host genome.[10]
dsDNA viruses can be subdivided between those that replicate in the nucleus, and as such are relatively dependent on host cell machinery for transcription and replication, and those that replicate in the cytoplasm, in which case they have evolved or acquired their own means of executing transcription and replication.[3] dsDNA viruses are also commonly divided between tailed dsDNA viruses, referring to members of the realm Duplodnaviria, usually the tailed bacteriophages of the order Caudovirales, and tailless or non-tailed dsDNA viruses of the realm Varidnaviria.[11][12]
dsDNA viruses are classified into three of the four realms and include many taxa that are unassigned to a realm:
- All viruses in Adnaviria are dsDNA viruses. Viruses in this realm infect archaea.[13]
- All viruses in Duplodnaviria are dsDNA viruses. Viruses in this realm belong to two groups: tailed bacteriophages in Caudovirales and herpesviruses in Herpesvirales.[11]
- In Monodnaviria, members of the class Papovaviricetes are dsDNA viruses. Viruses in Papovaviricetes constitute two groups: papillomaviruses and polyomaviruses.[14]
- All viruses in Varidnaviria are dsDNA viruses. Viruses in this realm include adenoviruses, giant viruses, and poxviruses.[12]
- The following taxa that are unassigned to a realm exclusively contain dsDNA viruses:[12]
- Classes: Naldaviricetes[15]
- Families: Ampullaviridae, Bicaudaviridae, Clavaviridae, Fuselloviridae, Globuloviridae, Guttaviridae, Halspiviridae, Ovaliviridae, Plasmaviridae, Polydnaviridae, Portogloboviridae, Thaspiviridae
- Genera: Dinodnavirus, Rhizidiovirus
Group II: single-stranded DNA viruses
[edit]
The second Baltimore group contains viruses that have a single-stranded DNA (ssDNA) genome. ssDNA viruses have the same manner of transcription as dsDNA viruses. Because the genome is single-stranded, however, it is first made into a double-stranded form by a DNA polymerase upon entering a host cell. mRNA is then synthesized from the double-stranded form. The double-stranded form of ssDNA viruses may be produced either directly after entry into a cell or as a consequence of replication of the viral genome.[16][17] Eukaryotic ssDNA viruses are replicated in the nucleus.[3][18]
Most ssDNA viruses contain circular genomes that are replicated via rolling circle replication (RCR). ssDNA RCR is initiated by an endonuclease that bonds to and cleaves the positive strand, allowing a DNA polymerase to use the negative strand as a template for replication. Replication progresses in a loop around the genome by means of extending the 3′-end of the positive strand, displacing the prior positive strand, and the endonuclease cleaves the positive strand again to create a standalone genome that is ligated into a circular loop. The new ssDNA may be packaged into virions or replicated by a DNA polymerase to form a double-stranded form for transcription or continuation of the replication cycle.[16][19]
Parvoviruses contain linear ssDNA genomes that are replicated via rolling hairpin replication (RHR), which is similar to RCR. Parvovirus genomes have hairpin loops at each end of the genome that repeatedly unfold and refold during replication to change the direction of DNA synthesis to move back and forth along the genome, producing numerous copies of the genome in a continuous process. Individual genomes are then excised from this molecule by the viral endonuclease. For parvoviruses, either the positive or negative sense strand may be packaged into capsids, varying from virus to virus.[19][20]
Nearly all ssDNA viruses have positive sense genomes, but a few exceptions and peculiarities exist. The family Anelloviridae is the only ssDNA family whose members have negative sense genomes, which are circular.[18] Parvoviruses, as previously mentioned, may package either the positive or negative sense strand into virions.[17] Lastly, bidnaviruses package both the positive and negative linear strands.[18][21] In any case, the sense of ssDNA viruses, unlike for ssRNA viruses, is not sufficient to separate ssDNA viruses into two groups since all ssDNA viral genomes are converted to dsDNA forms prior to transcription and replication.[2]
ssDNA viruses are classified into one of the four realms and include several families that are unassigned to a realm:
- In Monodnaviria, all members except viruses in Papovaviricetes are ssDNA viruses.[14]
- The unassigned families Anelloviridae and Spiraviridae are ssDNA virus families.[14]
- Viruses in the family Finnlakeviridae contain ssDNA genomes. Finnlakeviridae is unassigned to a realm but is a proposed member of Varidnaviria.[12]
RNA viruses
[edit]RNA viruses have genomes made of ribonucleic acid (RNA) and comprise three groups: double-stranded RNA (dsRNA) viruses, positive sense single-stranded RNA (+ssRNA) viruses, and negative sense single-stranded RNA (-ssRNA) viruses. The majority of RNA viruses are classified in the kingdom Orthornavirae in the realm Riboviria. The exceptions are generally viroids and other subviral agents. Some of the latter category, such as the hepatitis D virus, are classified in the realm Ribozyviria.
Group III: double-stranded RNA viruses
[edit]
The third Baltimore group contains viruses that have a double-stranded RNA (dsRNA) genome. After entering a host cell, the dsRNA genome is transcribed to mRNA from the negative strand by the viral RNA-dependent RNA polymerase (RdRp). The mRNA may be used for translation or replication. Single-stranded mRNA is replicated to form the dsRNA genome. The 5′-end of the genome may be naked, capped, or covalently bound to a viral protein.[22][23]
dsRNA is not a molecule made by cells, so cellular life has evolved antiviral systems to detect and inactivate viral dsRNA. To counteract this, many dsRNA genomes are constructed inside of capsids, thereby avoiding detection inside of the host cell's cytoplasm. mRNA is forced out from the capsid in order to be translated or to be translocated from a mature capsid to a progeny capsid.[22][23][24] While dsRNA viruses typically have capsids, viruses in the families Amalgaviridae and Endornaviridae have not been observed to form virions and as such apparently lack capsids. Endornaviruses are also unusual in that unlike other RNA viruses, they possess a single, long open reading frame (ORF), or translatable portion, and a site-specific nick in the 5′ region of the positive strand.[24]
dsRNA viruses are classified into two phyla within the kingdom Orthornavirae of the realm Riboviria:[25]
- All viruses in Duplornaviricota are dsRNA viruses.
- In Pisuviricota, all members of the class Duplopiviricetes are dsRNA viruses.
Group IV: positive sense single-stranded RNA viruses
[edit]
The fourth Baltimore group contains viruses that have a positive sense single-stranded RNA (+ssRNA) genome. For +ssRNA viruses, the genome functions as mRNA, so no transcription is required for translation. +ssRNA viruses will also, however, produce positive sense copies of the genome from negative sense strands of an intermediate dsRNA genome. This acts as both a transcription and a replication process since the replicated RNA is also mRNA. The 5′-end may be naked, capped, or covalently bound to a viral protein, and the 3′-end may be naked or polyadenylated.[26][27][28]
Many +ssRNA viruses are able to have only a portion of their genome transcribed. Typically, subgenomic RNA (sgRNA) strands are used for translation of structural and movement proteins needed during intermediate and late stages of infection. sgRNA transcription may occur by commencing RNA synthesis within the genome rather than from the 5′-end, by stopping RNA synthesis at specific sequences in the genome, or by, as a part of both prior methods, synthesizing leader sequences from the viral RNA that are then attached to sgRNA strands. Because replication is required for sgRNA synthesis, RdRp is always translated first.[27][28][29]
Because the process of replicating the viral genome produces intermediate dsRNA molecules, +ssRNA viruses can be targeted by the host cell's immune system. To avoid detection, +ssRNA viruses replicate in membrane-associated vesicles that are used as replication factories. From there, only viral +ssRNA, which may be mRNA, enters the main cytoplasmic area of the cell.[26][27]
+ssRNA viruses can be subdivided between those that have polycistronic mRNA, which encodes a polyprotein that is cleaved to form multiple mature proteins, and those that produce subgenomic mRNAs and therefore undergo two or more rounds of translation.[3][30] +ssRNA viruses are included in three phyla in the kingdom Orthornavirae in the realm Riboviria:[25]
- All viruses in Lenarviricota are +ssRNA viruses.
- All viruses in Pisuviricota are +ssRNA viruses, excluding the class Duplopiviricetes, whose members have dsRNA genomes.
- All viruses in Kitrinoviricota are +ssRNA viruses.
Group V: negative sense single-stranded RNA viruses
[edit]The fifth Baltimore group contains viruses that have a negative sense, single-stranded RNA (-ssRNA) genome. mRNA, which is positive sense, is transcribed directly from the negative sense genome. The first process for -ssRNA transcription involves RdRp binding to a leader sequence on the 3′ end of the genome, transcribing a 5′ triphosphate-leader RNA that is capped, then stopping and restarting on a transcription signal which is capped, continuing until a stop signal is reached.[31] The second manner is similar but instead of synthesizing a cap, RdRp may make use of cap snatching, whereby a short sequence of host cell mRNA is taken and used as the 5′ cap of the viral mRNA.[32] Genomic -ssRNA is replicated from the positive sense antigenome in a similar manner as transcription, except in reverse using the antigenome as a template for the genome. RdRp moves from the 3′-end to the 5′-end of the antigenome and ignores all transcription signals when synthesizing genomic -ssRNA.[23][33]
Various -ssRNA viruses use special mechanisms for transcription. The manner of producing the polyA tail may be via polymerase stuttering, during which RdRp transcribes an adenine from uracil and then moves back in the RNA sequence with the mRNA to transcribe it again, continuing this process numerous times until hundreds of adenines have been added to the 3′-end of the mRNA.[34] Additionally, some -ssRNA viruses are ambisense, as both the positive and negative strands separately encode viral proteins, and these viruses produce two separate mRNA strands: one directly from the genome and one from a complementary strand.[35][36]
-ssRNA viruses can be subdivided informally between those that have nonsegmented and segmented genomes. Nonsegmented -ssRNA viruses replicate in the cytoplasm, and segmented -ssRNA viruses replicate in the nucleus. During transcription, the RdRp produces one monocistronic mRNA strand from each segment of the genome.[3][23][37] All -ssRNA viruses are classified in the phylum Negarnaviricota in the kingdom Orthornavirae in the realm Riboviria. Negarnaviricota only contains -ssRNA viruses, so "-ssRNA virus" is synonymous with Negarnaviricota.[25] Negarnaviricota is divided into two subphyla: Haploviricotina, whose members synthesize a cap structure on viral mRNA required for protein synthesis, and Polyploviricotina, whose members instead obtain caps on mRNA via cap snatching.[38]
Reverse transcribing viruses
[edit]Reverse transcribing (RT) viruses have genomes made of either DNA or RNA and replicate via reverse transcription. Two groups of reverse transcribing viruses exist: single-stranded RNA-RT (ssRNA-RT) viruses, and double-stranded DNA-RT (dsDNA-RT) viruses. Reverse transcribing viruses are classified in the kingdom Pararnavirae in the realm Riboviria.
Group VI: single-stranded RNA viruses with a DNA intermediate
[edit]The sixth Baltimore group contains viruses that have a (positive-sense) single-stranded RNA genome that has a DNA intermediate ((+)ssRNA-RT) in its replication cycle.[note 1] ssRNA-RT viruses are transcribed in the same manner as DNA viruses, but their linear genomes are first converted to a dsDNA form through a process called reverse transcription. The viral reverse transcriptase enzyme synthesizes a DNA strand from the ssRNA strand, and the RNA strand is degraded and replaced with a DNA strand to create a dsDNA genome. The genome is then integrated into the DNA of the host cell, where it is now called a provirus. The host cell's RNA polymerase II then transcribes RNA in the nucleus from the proviral DNA. Some of this RNA may become mRNA whereas other strands will become copies of the viral genome for replication.[37][39][40][41]
ssRNA-RT viruses are all included in the class Revtraviricetes, phylum Arterviricota, kingdom Pararnavirae of the realm Riboviria. Excluding Caulimoviridae, which belongs to Group VII, all members of the Revtraviricetes order Ortervirales are ssRNA-RT viruses.[25][42]
Group VII: double-stranded DNA viruses with an RNA intermediate
[edit]The seventh Baltimore group contains viruses that have a double-stranded DNA genome that has an RNA intermediate (dsDNA-RT) in its replication cycle. dsDNA-RT viruses have a gap in one strand, which is repaired to create a complete dsDNA genome prior to transcription.[3][37] dsDNA-RT viruses are transcribed in the same manner as dsDNA viruses,[2] but make use of reverse transcription to replicate their circular genome while it is still in the capsid. The host cell's RNA polymerase II transcribes RNA strands from the genome in the cytoplasm, and the genome is replicated from these RNA strands. The dsDNA genome is produced from pregenomic RNA strands via the same general mechanism as ssRNA-RT viruses, but with replication occurring in a loop around the circular genome. After replication, the dsDNA genome may be packed or sent to the nucleus for further rounds of transcription.[39][43]
dsDNA-RT viruses are, like ssRNA-RT, all included in the class Revtraviricetes. Two families of dsDNA-RT viruses are recognized: Caulimoviridae, which belongs to the order Ortervirales, and Hepadnaviridae, which is the sole family in the order Blubervirales.[25][42]
Multi-group characteristics
[edit]
A number of characteristics of viruses are not directly associated with Baltimore classification but nonetheless closely correspond to multiple, specific Baltimore groups. This includes alternative splicing during transcription, whether the viral genome is segmented, the host range of viruses, whether the genome is linear or circular, and different methods of translating viral mRNA.
Alternative splicing
[edit]Alternative splicing is a mechanism by which different proteins can be produced from a single gene by means of using alternative splicing sites to produce different mRNAs. It is found in various DNA, -ssRNA, and reverse transcribing viruses. Viruses may make use of alternative splicing solely to produce multiple proteins from a single pre-mRNA strand or for other specific purposes. For certain viruses, including the families Orthomyxoviridae and Papillomaviridae, alternative splicing acts as a way to regulate early and late gene expression during different stages of infection. Herpesviruses use it as a potential anti-host defense mechanism to prevent synthesis of specific antiviral proteins. Furthermore, in addition to alternative splicing, because cellular unspliced RNA cannot be transported out of the nucleus, hepadnaviruses and retroviruses contain their own proteins for exporting their unspliced genomic RNA out of the nucleus.[44][45]
Genome segmentation
[edit]Viral genomes can exist in a single, or monopartite, segment, or they may be split into more than one molecule, called multipartite. For monopartite viruses, all genes are on the single segment of the genome. Multipartite viruses typically package their genomes into a single virion so that the whole genome is in one virus particle, and the separate segments contain different genes. Monopartite viruses are found in all Baltimore groups, whereas multipartite viruses are usually RNA viruses. This is because most multipartite viruses infect plants or fungi, which are eukaryotes, and most eukaryotic viruses are RNA viruses.[46][47][48] The family Pleolipoviridae varies as some viruses are monopartite ssDNA while others are bipartite with one segment being ssDNA and the other dsDNA.[6][49] Viruses in the ssDNA plant virus family Geminiviridae likewise vary between being monopartite and bipartite.[47][50]
Host range
[edit]Different Baltimore groups tend to be found within different branches of cellular life. In prokaryotes, the large majority of viruses are dsDNA viruses, and a significant minority are ssDNA viruses. Prokaryotic RNA viruses, in contrast, are relatively rare. Most eukaryotic viruses, including most animal and plant viruses, are RNA viruses, although eukaryotic DNA viruses are also common. By group, the vast majority of dsDNA viruses infect prokaryotes, ssDNA viruses are found in all three domains of life, dsRNA and +ssRNA viruses are primarily found in eukaryotes but also in bacteria, and -ssRNA and reverse transcribing viruses are only found in eukaryotes.[47][46][51]
Linear vs circular genomes
[edit]Viral genomes may be either linear with ends or circular in a loop. Whether a virus has a linear or circular genome varies from group to group. A significant percentage of dsDNA viruses are both, ssDNA viruses are primarily circular, RNA viruses and ssRNA-RT viruses are typically linear, and dsDNA-RT viruses are typically circular.[52][53] In the dsDNA family Sphaerolipoviridae, and in the family Pleolipoviridae, viruses contain both linear and circular genomes, varying from genus to genus.[6][49][54]
RNA editing
[edit]RNA editing is used by various ssRNA viruses to produce different proteins from a single gene. This can be done via polymerase slippage during transcription or by post-transcriptional editing. In polymerase slippage, the RNA polymerase slips one nucleotide back during transcription, inserting a nucleotide not included in the template strand. Editing of a genomic template would impair gene expression, so RNA editing is only done during and after transcription. For ebola viruses, RNA editing improves the ability to adapt to their hosts.[45][55]
Alternative splicing differs from RNA editing in that alternative splicing does not change the mRNA sequence like RNA editing but instead changes the coding capacity of an mRNA sequence as a result of alternative splicing sites. The two mechanisms otherwise have the same result: multiple proteins are expressed from a single gene.[45]
Translation
[edit]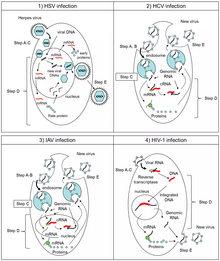
Translation is the process by which proteins are synthesized from mRNA by ribosomes. Baltimore groups do not directly pertain to the translation of viral proteins, but various atypical types of translation used by viruses are usually found within specific Baltimore groups:[2][56]
- Non-canonical translation initiation:
- Viral initiation of translation: used primarily by +ssRNA and ssRNA-RT viruses, various viruses have evolved mechanisms to initiate translation, such as having internal ribosomal entry sites to allow for cap-independent translation, having downstream hairpin loops that allow for cap-dependent translation in the absence of an eIF2 initiation factor, and initiation at a CUG or other start codon with a leucine amino acid.[57][58]
- Leaky scanning: used by various viruses in all Baltimore groups, the 40S ribosomal subunit may scan through a start codon, thereby skipping an ORF, only initiating translation with the 60S subunit at a subsequent start codon.[59][60]
- Ribosomal shunting: used by various dsDNA, +ssRNA, -ssRNA, ssRNA-RT, a dsDNA-RT viruses, ribosomes will start scanning from a 5′-cap structure then bypass a leader structure in the mRNA, initiation translation downstream from the leader sequence.[61][62]
- Termination-reinitiation: used by some dsRNA and +ssRNA viruses, ribosomes may translate an ORF, but following termination of translation of that ORF, a proportion of 40S subunits of the ribosome remain attached to the mRNA as a way to reinitiate translation of a subsequent ORF.[63]
- Non-canonical elongation and termination of translation:
- Ribosomal frameshifting: used by various dsDNA, dsRNA, +ssRNA, and ssRNA-RT viruses, produces merged proteins from overlapping ORFs. This is executed simply by ribosomes slipping one nucleobase forward or backward during translation.[60][64]
- Suppression of termination: also called stop-codon readthrough, used by various dsRNA, +ssRNA, and ssRNA-RT viruses, certain viruses contain codons in their mRNA that would normally signal for termination of translation upon being recognized by a release factor but are instead partially recognized by tRNA during translation, which allows for continued translation up to the next stop codon in order to produce an extended end of the viral protein.[65] In viruses, this is often used to express replicase enzymes.[66]
- Ribosomal skipping: also called stop-carry on, used by various dsRNA and +ssRNA viruses, a viral peptide, or amino acid sequence, may prevent a ribosome from covalently linking a new inserted amino acid, which blocks further translation. Consequently, the polyprotein is co-translationally cleaved, and a new amino acid sequence is started, leading to the production of two individual proteins from one ORF.[62][67]
History
[edit]
Baltimore classification was proposed in 1971 by virologist David Baltimore in a paper titled Expression of Animal Virus Genomes. It initially contained the first six groups but was later expanded to include group VII.[37][68][69] Because of the utility of Baltimore classification, it has come to be used alongside standard virus taxonomy, which is based on evolutionary relationships and governed by the International Committee on Taxonomy of Viruses (ICTV).[69]
From the 1990s to the 2010s, virus taxonomy used a 5-rank system ranging from order to species with Baltimore classification used in conjunction. Outside of the ICTV's official framework, various supergroups of viruses joining different families and orders were created over time based on increasing evidence of deeper evolutionary relations. Consequently, in 2016, the ICTV began to consider establishing ranks higher than order as well as how the Baltimore groups would be treated among higher taxa.[69]
In two votes in 2018 and 2019, a 15-rank system ranging from realm to species was established by the ICTV.[69] As part of this, the Baltimore groups for RNA viruses and RT viruses were incorporated into formal taxa. In 2018, the realm Riboviria was established and initially included the three RNA virus groups.[70] A year later, Riboviria was expanded to also include both RT groups. Within the realm, RT viruses are included in the kingdom Pararnavirae and RNA viruses in the kingdom Orthornavirae. Furthermore, the three Baltimore groups for RNA viruses are used as defining characteristics of the phyla in Orthornavirae.[25]
Unlike RNA viruses and RT viruses, DNA viruses have not been united under a single realm but are instead dispersed across four realms and various taxa that are not assigned to a realm. The realms Adnaviria and Duplodnaviria exclusively contains dsDNA viruses,[11][13] Monodnaviria primarily contains ssDNA viruses but also contains dsDNA viruses,[14] and Varidnaviria exclusively contains dsDNA viruses, although some proposed members of Varidnaviria, namely the family Finnlakeviridae, are ssDNA viruses.[12]
Explanatory notes
[edit]- ^ ssRNA-RT viruses are often called retroviruses, although this term is also used to refer to any reverse transcribing virus as well as specifically to viruses in the ssRNA-RT family Retroviridae.
References
[edit]Citations
[edit]- ^ a b Lostroh 2019, pp. 11–13.
- ^ a b c d "Viral replication/transcription/translation". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b c d e f Cann 2015, pp. 122–127.
- ^ "dsDNA templated transcription". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Rampersad & Tennant 2018, p. 66.
- ^ a b c Fermin 2018, pp. 36–40.
- ^ "dsDNA bidirectional replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "dsDNA rolling circle replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "DNA strand displacement replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Replicative transposition". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b c Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal/primary taxonomic ranks, for dsDNA viruses encoding HK97-type major capsid proteins" (docx). International Committee on Taxonomy of Viruses. Retrieved 6 August 2020.
- ^ a b c d e Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for DNA viruses encoding vertical jelly roll-type major capsid proteins" (docx). International Committee on Taxonomy of Viruses. Retrieved 6 August 2020.
- ^ a b Krupovic M, Kuhn JH, Wang F, Baquero DP, Egelman EH, Koonin EV, Prangishvili D (31 July 2020). "Create one new realm (Adnaviria) for classification of filamentous archaeal viruses with linear dsDNA genomes" (docx). International Committee on Taxonomy of Viruses (ICTV). Retrieved 27 May 2021.
- ^ a b c d Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for ssDNA viruses" (docx). International Committee on Taxonomy of Viruses. Retrieved 6 August 2020.
- ^ Harrison RL, Herniou EA, Jehle JA, Theilmann DA, Burand JP, Krell PJ, van Oers MM (28 August 2020). "Create one new class (Naldaviricetes)including one new order (Lefavirales) for four families of arthropod-specific large DNA viruses" (docx). International Committee on Taxonomy of Viruses (ICTV). Retrieved 27 May 2021.
- ^ a b "ssDNA Rolling circle". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b "Rolling hairpin replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b c Fermin 2018, pp. 40–41.
- ^ a b Rampersad & Tennant 2018, pp. 61–62.
- ^ Kerr J, Cotmore S, Bloom ME (25 November 2005). Parvoviruses. CRC Press. pp. 171–185. ISBN 9781444114782.
- ^ "Bidnaviridae". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b "Double-stranded RNA virus replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b c d Rampersad & Tennant 2018, p. 65.
- ^ a b Fermin 2018, p. 42.
- ^ a b c d e f Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for realm Riboviria" (docx). International Committee on Taxonomy of Viruses (ICTV). Retrieved 6 August 2020.
- ^ a b "Positive stranded RNA virus replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b c Rampersad & Tennant 2018, pp. 64–65.
- ^ a b Fermin 2018, pp. 43–44.
- ^ "Subgenomic RNA transcription". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Cann 2015, pp. 151–154.
- ^ "Negative-stranded RNA virus transcription". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Cap snatching". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Negative stranded RNA virus replication". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Negative-stranded RNA virus polymerase stuttering". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Ambisense transcription in negative stranded RNA viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Cann 2015, pp. 154–156.
- ^ a b c d Fermin 2018, pp. 45–46.
- ^ Kuhn JH, Wolf YI, Krupovic M, Zhang YZ, Maes P, Dolja VV, Koonin EV (February 2019). "Classify viruses - the gain is worth the pain" (PDF). Nature. 566 (7744): 318–320. Bibcode:2019Natur.566..318K. doi:10.1038/d41586-019-00599-8. PMID 30787460. S2CID 67769904. Retrieved 6 August 2020.
- ^ a b Rampersad & Tennant 2018, pp. 63–64.
- ^ "ssRNA(RT) replication/transcription". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Cann 2015, p. 156.
- ^ a b Krupovic M, Blomberg J, Coffin JM, Dasgupta I, Fan H, Geering AD, Gifford R, Harrach B, Hull R, Johnson W, Kreuze JF, Lindemann D, Llorens C, Lockhart B, Mayer J, Muller E, Olszewski NE, Pappu HR, Pooggin MM, Richert-Poggeler KR, Sabanadzovic S, Sanfacon H, Schoelz JE, Seal S, Stavolone L, Stoye JP, Teycheney PY, Tristem M, Koonin EV, Kuhn JH (15 June 2018). "Ortervirales: New Virus Order Unifying Five Families of Reverse-Transcribing Viruses". J Virol. 92 (12): e00515 – e00518. doi:10.1128/JVI.00515-18. PMC 5974489. PMID 29618642.
- ^ "dsDNA(RT) replication/transcription". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Alternative splicing". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b c Rampersad & Tennant 2018, pp. 71–72.
- ^ a b Koonin EV, Dolja VV, Krupovic M (May 2015). "Origins and Evolution of Viruses of Eukaryotes: The Ultimate Modularity". Virology. 479: 2–25. doi:10.1016/j.virol.2015.02.039. PMC 5898234. PMID 25771806.
- ^ a b c Fermin 2018, pp. 35–46.
- ^ Sicard A, Michalakis Y, Gutierrez S, Blanc S (3 November 2016). "The Strange Lifestyle of Multipartite Viruses". PLOS Pathog. 12 (11): e1005819. doi:10.1371/journal.ppat.1005819. PMC 5094692. PMID 27812219.
- ^ a b Bamford DH, Pietilä MK, Roine E, Atanasova NS, Dienstbier A, Oksanen HM (December 2017). "ICTV Virus Taxonomy Profile: Pleolipoviridae". J Gen Virol. 98 (12): 2916–2917. doi:10.1099/jgv.0.000972. PMC 5882103. PMID 29125455. Retrieved 6 August 2020.
- ^ "Geminiviridae". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Wolf YI, Kazlauskas D, Iranzo J, Lucia-Sanz A, Kuhn JH, Krupovic M, Dolja VV, Kooning EV (27 November 2018). "Origins and Evolution of the Global RNA Virome". mBio. 9 (6): e02329-18. doi:10.1128/mBio.02329-18. PMC 6282212. PMID 30482837.
- ^ Yu C, Hernandez T, Zheng H, Yau SC, Huang HH, He RL, Yang J, Yau SS (22 May 2013). "Real Time Classification of Viruses in 12 Dimensions". PLOS ONE. 8 (5): e64328. Bibcode:2013PLoSO...864328Y. doi:10.1371/journal.pone.0064328. PMC 3661469. PMID 23717598.
- ^ "Double strand DNA viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
"Single strand DNA viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
"Double strand RNA viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
"Positive strand RNA viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
"Negative strand RNA viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
"Reverse-transcribing viruses". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020. - ^ "Sphaerolipoviridae". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "RNA editing". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Firth AE, Brierley I (July 2012). "Non-canonical translation in RNA viruses". J Gen Virol. 9 (Pt 7): 1385–1409. doi:10.1099/vir.0.042499-0. PMC 3542737. PMID 22535777.
- ^ "Viral initiation of translation". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Rampersad & Tennant 2018, pp. 69–70.
- ^ "Leaky scanning". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b Rampersad & Tennant 2018, pp. 73–74.
- ^ "Ribosomal shunt". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ a b Rampersad & Tennant 2018, pp. 74–75.
- ^ "RNA termination-reinitiation". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "Ribosomal frameshifting". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ "RNA suppression of termination". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Rampersad & Tennant 2018, pp. 72–73.
- ^ "Ribosomal skipping". ViralZone. Swiss Institute of Bioinformatics. Retrieved 6 August 2020.
- ^ Baltimore D (1971). "Expression of animal virus genomes". Bacteriol Rev. 35 (3): 235–241. doi:10.1128/MMBR.35.3.235-241.1971. PMC 378387. PMID 4329869.
- ^ a b c d International Committee on Taxonomy of Viruses Executive Committee (May 2020). "The New Scope of Virus Taxonomy: Partitioning the Virosphere Into 15 Hierarchical Ranks". Nat Microbiol. 5 (5): 668–674. doi:10.1038/s41564-020-0709-x. PMC 7186216. PMID 32341570.
- ^ Gorbalenya, Alexander E.; Krupovic, Mart; Siddell, Stuart; Varsani, Arvind; Kuhn, Jens H. (15 October 2018). "Riboviria: establishing a single taxon that comprises RNA viruses at the basal rank of virus taxonomy" (docx). International Committee on Taxonomy of Viruses (ICTV). Retrieved 6 August 2020.
General bibliography
[edit]- Cann, A. (2015). Principles of Molecular Virology. Elsevier. pp. 122–127. ISBN 978-0128019559.
- Fermin, G. (2018). Viruses: Molecular Biology, Host Interactions and Applications to Biotechnology. Elsevier. pp. 35–46. doi:10.1016/B978-0-12-811257-1.00002-4. ISBN 978-0128112571. S2CID 89706800.
- Lostroh, P. (2019). Molecular and Cellular Biology of Viruses. Garland Science. ISBN 978-0429664304.
- Rampersad, S.; Tennant, P. (2018). Viruses: Molecular Biology, Host Interactions and Applications to Biotechnology. Elsevier. pp. 55–82. doi:10.1016/B978-0-12-811257-1.00003-6. ISBN 978-0128112571. S2CID 90170103.
External links
[edit] Media related to Baltimore classification at Wikimedia Commons
Media related to Baltimore classification at Wikimedia Commons
